
George Shields
Professor, Chemistry
- Email: george.shields@furman.edu
- Phone: 864.294.2008
- Office: Townes Science Center 071F
George Shields enjoys a national reputation in the field of undergraduate research, having collaborated with more than 140 undergraduate students in the fields of computational chemistry, structural biochemistry and science education. He received the 2015 American Chemical Society (ACS) Award for Research at an Undergraduate Institution and the 2018 Research Corporation for Science Advancement Transformational Research and Excellence in Education Award (TREE). He was elected as a Fellow of the American Association for the Advancement of Science (AAAS) in 2019, and as a Fellow of the American Chemical Society (ACS) in 2020. He received the Council of Undergraduate Research (CUR) Fellow Award in 2020 and the CUR-Goldwater Scholars Faculty Mentor Award in 2022. He is the founder and director of the Molecular Education and Research Consortium in Undergraduate Computational Chemistry (MERCURY), a collaboration of 45 undergraduate research teams at 41 different institutions. He serves on the advisory board of the Journal of Physical Chemistry.
George’s students have received 42 national awards, including four Fulbright, 14 Goldwater, and eight graduate fellowships, and 90% of his alumni have pursued advanced degrees in science, medicine, law, or business. Since 1993, he has published 114 scientific research papers, including 74 papers with 69 undergraduate co-authors. Eighteen of his papers have been cited more than a hundred times (Web of Science), including three papers in Science, five papers in the Journal of the American Chemical Society, and five papers in the Journal of Physical Chemistry. His undergraduates have presented at more than 200 national and international conferences. He has received over $6.3 million in funding to support undergraduate research.
George served as Furman's Vice President for Academic Affairs and Provost from 2016-2019. This was preceded by six years at Bucknell University as Dean of the College of Arts and Sciences and professor in the department of chemistry. He served as founding Dean of the College of Science and Technology at Armstrong Atlantic State University, and he also taught in and chaired the chemistry departments at Hamilton College and Lake Forest College.
Education
- Ph.D., Georgia Institute of Technology
- M.S., Georgia Institute of Technology
- B.S., Georgia Institute of Technology
Research
My research efforts use computational methods to gain insights into biochemistry and atmospheric chemistry. My research group uses quantum chemistry, Monte Carlo, and molecular dynamics techniques to investigate the structure and function of molecules. The common theme throughout is the involvement of undergraduates in a productive and meaningful research experience.
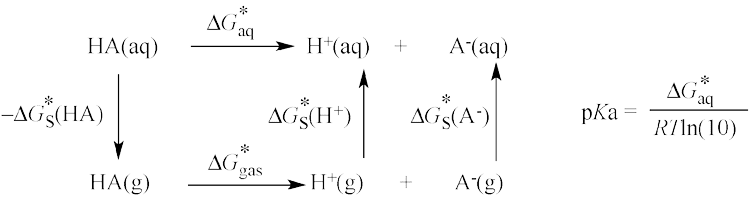
Computational pKa Prediction
Using these methods requires a thorough understanding of solvation effects, and much of our basic work involves finding and learning how to use the best methods for incorporating solvation into traditional computational chemistry techniques. We have completed a systematic study of pKa calculations, funded by the American Chemistry Society (ACS), Petroleum Research Fund (PRF) and National Science Foundation (NSF), in order to learn how state-of-the-art methods can best be used to accurately predict deprotonation in aqueous solution.
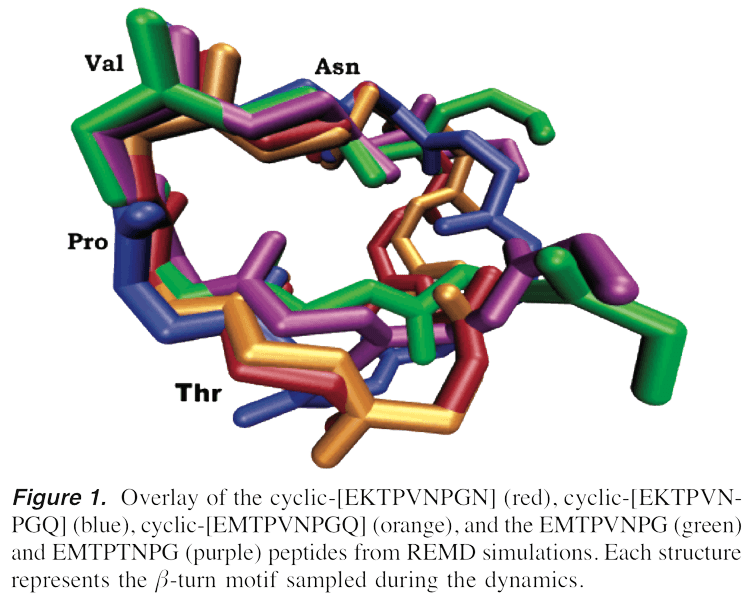
Drug Design
We had drug design projects with funding from NIH, DOD, and Research Corporation, with an emphasis on breast cancer. Alpha-fetoprotein (AFP) is a protein produced by the fetal yolk sac and is presumed to act as a growth regulator during gestation. It is 591 amino acids long. It was discovered that high levels of circulating AFP in maternal serum decrease a woman’s risk of developing estrogen receptor positive (ER+) breast cancer later in life. It has recently been discovered that the peptide can be shortened to peptides as small as 4 amino acids long and still retain active breast cancer inhibition qualities. We have applied and demonstrated for the first time that replica exchange molecular dynamics (REMD) simulations can be used as a novel lead compound design tool. We have shown that a common conformation that is shared between the active linear 8-mer and cyclic 9-mer peptides of AFP is a conserved reverse beta-turn, and the smaller peptide analogs TOVNO, TPVNP, TOVN, and TPVN. These analogs inhibit estrogen-dependent cell growth in a mouse uterine growth assay, through interaction with a yet to be discovered key receptor, and inhibit human breast cancer in a mouse xenograft.
Structure and Properties of Water Clusters and Atmospheric Hydrates
We are also working on projects in atmospheric chemistry that focus on the role of water clusters. The aim is to locate the global and all relevant low lying local minima for each cluster.
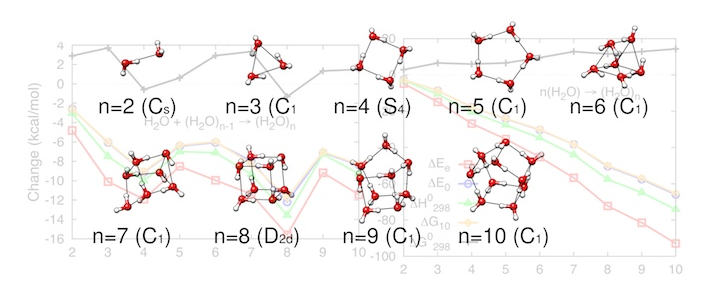
Our sampling methods ensure that we have searched the large configurational space of hydrogen bonding networks available to these clusters and extracted the lowest energy configurations. Our quantum mechanical method of choice, MP2 is the most affordable yet very accurate method for studying non-covalently bonded systems like water clusters. The interplay between energy and entropy, the shapes and hydrogen bonding networks of the clusters are used to explain the growth patterns of the clusters. Ultimately, this information will be used to calculate cluster populations and nucleation rates that are compared with experiment rates of formation of aerosol particles.
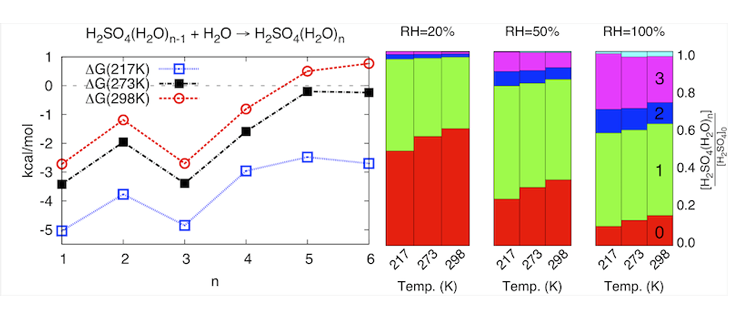
Improved understanding of atmospheric aerosol formation would reduce the large uncertainty in the cooling effect of aerosols on the global radiation balance (see below) and refine global climate models.
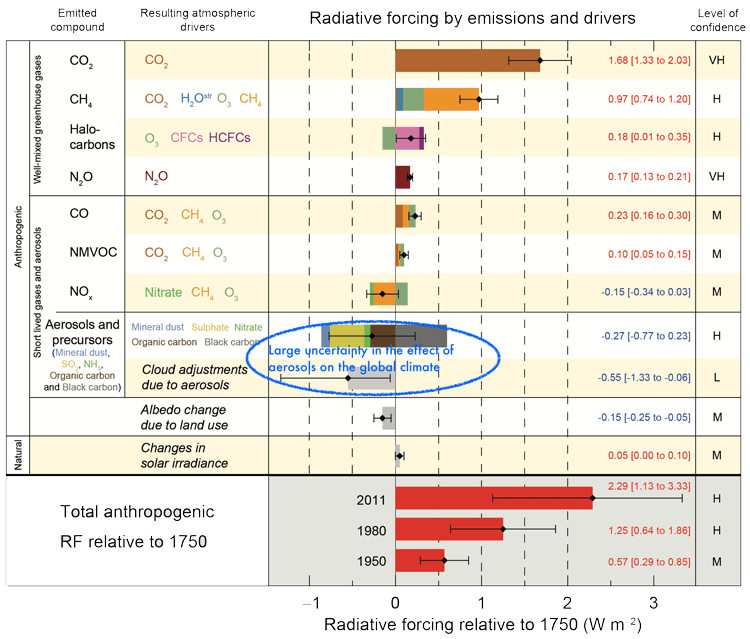
IPCC WG1 4th Assessment Report, 2007
-
Longsworth, O.M.; Bready, C.J.; Shields, G.C. "The Driving Effects of Common Atmospheric Molecules for Formation of Clusters: The Case of Sulfuric Acid, Formic Acid, Hydrochloric Acid, Ammonia, and Dimethylamine" Environ. Sci.: Atmos. 2023, 3, 1335-1351
-
Ball, N.D.; Gomez, M.A.; Rempel, B.P.; Farkas, E.R.; Makal, T.E.; Shields, G.C.; Parish, C.A.; Tresc, B.W.; McGinitie, E.G.; Edwards, E. “Conducting research at primarily undergraduate institutions” Cell Rep. Phys. Sci. (2023), 4, 1-6.
-
Elm, J.; Ayoubi, D.; Engsvang, M.; Jensen, A.B.; Knattrup, Y.; Kubecka, J.; Bready, C.J.; Fowler, V.R.; Harold, S.E.; Longsworth, O.M; Shields, G.C. "Quantum chemical modeling of organic enhanced atmospheric nucleation: A critical review." WIREs Comput. Mol. Sci. (2023), e1662.
-
Longsworth, O. M.; Bready, C. J.; Joines, M. S.; Shields, G. C. "The Driving Effects of Common Atmospheric Molecules for Formation of Prenucleation Clusters: The Case of Sulfuric Acid, Nitric Acid, Hydrochloric Acid, Ammonia, and Dimethyl Amine", Environ. Sci. Atmos. 2023, 3, 1585 – 1600.
-
Harold, S. E. ; Warf, S. L. ; Shields, G. C. “Prebiotic Dimer and Trimer Peptide Formation in Gas-Phase Atmospheric Nanoclusters of Water”, Phys. Chem. Chem. Phys., 2023, 25, 28517 – 28532.
-
Bready, C.J.; , Vanovac, S.; Odbadrakh, T.T.; Shields, G.C.; “Amino Acids Compete with Ammonia in Sulfuric Acid-Based Atmospheric Aerosol Prenucleation: The Case of Glycine and Serine.” J. Phys. Chem. A 2022 126. 5195-5206.
-
Afzalifar, A.; Shields, G.C.; Fowler, V.R; Ras, R. “Probing Free Energy of Small Water Clusters: Revisiting Classical Nucleation Theory." J. Phys. Chem. Lett. 2022 13 8038-8046.
-
London, H.C.; Pritchett, D.Y.; Pienkos, J.A.; McMillen, C.D.; Whittemore, T.J.; Bready, C.J.; Myers, A.R.; Vieira, N.C.; Harold, S.; Shields, G.C.; Wagenknecht. P.S. "Photochemistry and Photophysics of Charge-Transfer Excited States in Emissive d10/d0 Heterobimetallic Titanocene Tweezer Complexes." Inorg. Chem. 2022, 61, 28, 10986–10998.
-
Bready, C.J.; Fowler, V.R.; Juechter, L.A.; Kurfman, L.A.; Mazaleski, G.E.; Shields, G.C. "The Driving Effects of Common Atmospheric Molecules for Formation of Prenucleation Clusters: The Case of Sulfuric Acid, Formic Acid, Nitric Acid, Ammonia, and Dimethyl Amine" Environ. Sci.: Atmos. 2022 , 2, 1469-1486.
-
Harold, S.E.; Bready, C.J.; Juechter, L.A.; Kurfman, L.A.; Vanovac, S.; Fowler, V.R.; Mazaleski, G.E.; Odbadrakh, T.T.; Shields, G.C. “Hydrogen-Bond Topology is More Important than Acid/Base Strength in Atmospheric Prenucleation Clusters.” J. Phys. Chem. A 2022, 126, 10, 1718-1728.
-
Shields, G.C. “Physical Chemistry Research in the Shields Lab by Goldwater Scholars: Including Lessons Learned” in Physical Chemistry Research at Undergraduate Institutions: Innovative and Impactful Approaches, Volume 2. Todd A. Hopkins, Carol A. Parish, editors, ACS, Washington, D.C., ACS Symposium Series, Vol 1429, Chapter 1, 2022, 1-30.
-
Calabrese, C.; Temelso, B.; Usabiaga, I.; Seifert, N.A.; Basterretxea, F.J.; Prampolini, G.; Shields, G.C.; Pate, B.H.; Evangelisti, L.; Cocinero, E.J. "The Role of Non-Covalent Interactions on Cluster Formation: Pentamer, Hexamers and Heptamer of Difluoromethane." Angew. Chem. Int. Ed., 2021, 60, 16894.
-
Ball, B.T.; Vanovac, S.; Odbadrakh, T.T; Shields, G.C. "Monomers of Glycine and Serine Have a Limited Ability to Hydrate in the Atmosphere." J. Phys. Chem. A 2021, 125, 38, 8454-8467
-
Carlton, E. S.; Sutton, J. J.; Gale, A. G.; Shields, G. C.; Gordon, K. C.; Wagenknecht, P. S. "Insights into the Charge-Transfer Character of Electronic Transitions in RCp2Ti(C2Fc)2 Complexes using Solvatochromism, Resonance Raman Spectroscopy, and TDDFT" Dalton Trans. 2021, 50, 2233-2242.
-
London, H.C.; Whittemore, T.J.; Gale, A.G.; McMillen, C.D.; Pritchett, D.Y.; Myers, A.R.; Thomas, H.D.; Shields, G.C.; Wagenknecht, P.S. "Ligand-to-Metal Charge-Transfer Photophysics and Photochemistry of Emissive d0 Titanocenes: A Spectroscopic and Computational Investigation." Inorg. Chem., 2021, 60, 14399-14409.
-
Kurfman, L.A.; Odbadrakh, T.T.; Shields, G.C. "Calculating Reliable Gibbs Free Energies for Formation of Gas-Phase Clusters that Are Critical for Atmospheric Chemistry: (H2SO4)3", J. Phys. Chem. A 2021, 125, 15, 3169-3176.
-
Gale, A.G., Odbadrakh, T.T., Ball, B.T., and Shields, G.C. "Water-Mediated Peptide Bond Formation in the Gas Phase: A Model Prebiotic Reaction", J. Phys. Chem. A
, 2020, 124 (20), 4150-4159. -
Leonardi, A., Ricker, H.M., Gale, A.G., Ball, B.T., Odbadrakh, T.T., Shields, G.C., Navea, J.G. "Particle formation and surface processes on atmospheric aerosols: A review of applied quantum chemical calculations." Int J Quantum Chem., 2020, e26350.
-
Shields, G.C., Feller, S.E. "Maintaining a high degree of research productivity at a predominately undergraduate institution as your career advances" Int J Quantum Chem., 2020, e26370.
-
Odbadrakh, T.T., Gale, A.G., Ball, B.T., Temelso, B., Shields, G.C. "Computation of Atmospheric Concentrations of Molecular Clusters from ab initio Thermochemistry" J. Vis. Exp., 2020, 158.
-
Gale A.G., Odbadrakh T.T., Shields G.C.. "Catalytic activity of water molecules in gas-phase glycine dimerization", Int J Quantum Chem., 2020, e26469.
-
Shields, G.C. "Twenty years of exceptional success: The Molecular Education and Research Consortium in Undergraduate Computational Chemistry (MERCURY)." Int J Quantum Chem., 2020, e26274.
-
Shields, G.C. The Molecular Education and Research Consortium in Computational Chemistry: Twenty Years of Exceptional Success Supporting Undergraduate Research and Inclusive Excellence. SPUR, 2019, 3, 5-15.
-
López, J.C.; Pérez, C.; Blanco, S.; Shubert, V.A; Temelso, B.; Shields, G.C.; Schnell, M. Water Induces the Same Crown Shapes as Li+ or Na+ in 15-crown-5 Ether: a Broadband Rotational Study. Phys. Chem. Chem. Phys. 2019, 21, 2875-2881.
-
Temelso, B.; Morrison, E . F.; Speer, D. L.; Cao, B. C.; Appiah-Padi, N.; Kim, G.; Shields, G. C. Effect of Mixing Ammonia and Alkylamines on Sulfate Aerosol Formation. J. Phys. Chem. A, 2018, 122, 1612−1622
-
Temelso, B.; Klein K. L.; Mabey, J. W.; Perez, C.; Pate, B. H.; Kisiel, Z.; Shields, G. C. Exploring the Rich Potential Energy Surface of (H2O)11 and Its Physical Implications J. Chem. Theory Comput., 2018, 14, 1141-1153.
-
Pérez, C.; Steber, A. L.; Rijs, A. M.; Temelso, B.; Shields, G. C.; Lopez, J. C.; Kisiel, Z.; Schnell, M. Corannulene and its complex with water: a tiny cup of water. Phys. Chem. Chem. Phys. 2017, 19, 14214-14223.
-
Temelso, B.; Mabey, J. M.; Kubota, T.; Appiah-Padi, N.; Shields, G. C. ArbAlign: A Tool for Optimal Alignment of Arbitrarily Ordered Isomers Using the Kuhn-Munkres Algorithm. J. Chem. Inf. Model., 2017, 57, 1045-1054.
-
Steber, A. L.; Pérez, C.; Temelso, B.; Shields, G. C.; Rijs, A. M.; Pate, B. H.; Kisiel, Z.; Schnell, M. Capturing the Elusive Water Trimer from the Stepwise Growth of Water on the Surface of the Polycyclic Aromatic Hydrocarbon Acenaphthene. J. Phys. Chem. Lett., 2017, 8, 5744-5750.
-
Richardson, J. O.; Pérez, C.; Lobsiger, S.; Reid, A. A.; Temelso, B.; Shields, G. C.; Kisiel, Z.; Wales, D. J.; Pate, B. H.; Althorpe, S. C. Concerted Hydrogen-Bond Breaking by Quantum Tunneling in the Water Hexamer Prism. Science. 2016, 351 (6279), 1310-1313.
-
Shields, G. C. Guest Foreword. In The Power and Promise of Early Research; Murray, D. H., Obare, S. O., Hageman, J. H., Eds.; American Chemical Society: Washington, D.C., 2016; pp xiii-xv.
-
Swan, J. S.; Findeis, P. M.; Hilton, S.; Lebold, K. M.; Temelso, B.; Shields, G. C. Formation of Deprotonated 2-Imidazoline-4(5)-One Product Ions in the Collision-Induced Dissociation of Some Serine-Containing Dipeptides. Int. J. Mass Spectrom. 2015, 381, 25-32.
-
Temelso, B.; Renner, C. R.; Shields, G. C. Importance and Reliability of Small Basis Set CCSD(T) Corrections to MP2 Binding and Relative Energies of Water Clusters. J. Chem. Theory Comput. 2015, 11 (4), 1439-1448.
-
Seybold, P. G.; Shields, G. C. Computational Estimation of pKa Values. Wiley Interdiscip. Rev. Comput. Mol. Sci. 2015, 5 (3), 290-297.
-
Bustos, D. J.; Temelso, B.; Shields, G. C. Hydration of the Sulfuric Acid-Methylamine Complex and Implications for Aerosol Formation. J. Phys. Chem. A 2014, 118 (35), 7430-7441.
-
Pérez, C.; Zaleski, D. P.; Seifert, N. A.; Temelso, B.; Shields, G. C.; Kisiel, Z.; Pate, B. H. Hydrogen Bond Cooperativity and the Three-Dimensional Structures of Water Nonamers and Decamers. Angew. Chemie Int. Ed. 2014, 53 (52), 14368-14372.
-
Temelso, B.; Alser, K. A.; Gauthier, A.; Palmer, A. K.; Shields, G. C. Structural Analysis of α-Fetoprotein (AFP)-like Peptides with Anti-Breast-Cancer Properties. J. Phys. Chem. B 2014, 118 (17), 4514-4526.
-
Shields, G. C.; Seybold, P. G. Computational Approaches for the Prediction of pKa Values; CRC Press: Boca Raton, 2014.
-
Pérez, C.; Lobsiger, S.; Seifert, N. A.; Zaleski, D. P.; Temelso, B.; Shields, G. C.; Kisiel, Z.; Pate, B. H. Broadband Fourier Transform Rotational Spectroscopy for Structure Determination: The Water Heptamer. Chem. Phys. Lett. 2013, 571, 1-15.
-
Temelso, B.; Köddermann, T.; Kirschner, K. N.; Klein, K.; Shields, G. C. Structure and Thermodynamics of H3O+(H2O)8 Clusters: A Combined Molecular Dynamics and Quantum Mechanics Approach. Comput. Theor. Chem. 2013, 1021, 240-248.
-
Kinnel, R. B.; Van Wynsberghe, A. W.; Rosenstein, I. J.; Brewer, K. S.; Cotten, M.; Shields, G. C.; Borton, C. J.; Senior, S. Z.; Rahn, G. S.; Elgren, T. E. A Departmental Focus on High Impact Undergraduate Research Experiences. In Developing and Maintaining a Successful Undergraduate Research Program; American Chemical Society: Washington, D.C., 2013; pp 5-22.
-
Husar, D. E.; Temelso, B.; Ashworth, A. L.; Shields, G. C. Hydration of the Bisulfate Ion: Atmospheric Implications. J. Phys. Chem. A 2012, 116 (21), 5151-5163.
-
Pérez, C.; Muckle, M. T.; Zaleski, D. P.; Seifert, N. A.; Temelso, B.; Shields, G. C.; Kisiel, Z.; Pate, B. H. Structures of Cage, Prism, and Book Isomers of Water Hexamer from Broadband Rotational Spectroscopy. Science. 2012, 336 (6083), 897-901.
-
Temelso, B.; Morrell, T. E.; Shields, R. M.; Allodi, M. A.; Wood, E. K.; Kirschner, K. N.; Castonguay, T. C.; Archer, K. A.; Shields, G. C. Quantum Mechanical Study of Sulfuric Acid Hydration: Atmospheric Implications. J. Phys. Chem. A 2012, 116 (9), 2209-2224.
-
Temelso, B.; Phan, T. N.; Shields, G. C. Computational Study of the Hydration of Sulfuric Acid Dimers: Implications for Acid Dissociation and Aerosol Formation. J. Phys. Chem. A 2012, 116 (39), 9745-9758.
-
Temelso, B.; Archer, K. A.; Shields, G. C. Benchmark Structures and Binding Energies of Small Water Clusters with Anharmonicity Corrections. J. Phys. Chem. A 2011, 115 (43), 12034-12046.
-
Temelso, B.; Shields, G. C. The Role of Anharmonicity in Hydrogen-Bonded Systems: The Case of Water Clusters. J. Chem. Theory Comput. 2011, 7 (9), 2804-2817.
-
Andersen, T. T.; Bennett, J. A.; Jacobson, H.; Shields, G. C. Alpha-Fetoprotein Peptides. US 7964701 B2, 2011.
-
Shields, R. M.; Temelso, B.; Archer, K. A.; Morrell, T. E.; Shields, G. C. Accurate Predictions of Water Cluster Formation, (H20)n=2-10. J. Phys. Chem. A 2010, 114 (43), 11725-11737.
-
Alongi, K. S.; Shields, G. C. Theoretical Calculations of Acid Dissociation Constants: A Review Article. In Annual Reports in Computational Chemistry; Ralph A. Wheeler, Ed.; Elsevier: Amsterdam, 2010; Vol. 6, pp 113-138.
-
Morrell, T. E.; Shields, G. C. Atmospheric Implications for Formation of Clusters of Ammonium and 1-10 Water Molecules. J. Phys. Chem. A 2010, 114 (12), 4266-4271.
-
Shields, G. C. Creating a Comprehensive Summer Undergraduate Research Program Despite Fiscal Challenges. CUR Q. 2010, 30 (4), 18-21.
-
Shields, G. C. Using Pre-College Research to Promote Student Success and Increase the Number of Science Majors. CUR Q. 2010, 31 (1), 43-47.
-
Andersen, T. T.; Bennett, J. A.; Jacobson, H.; Shields, G. C.; Kirschner, K. N. Alpha-Fetoprotein Peptides and Uses Thereof. US 7598342 B2, 2009.
-
Joseph, L. C.; Bennett, J. A.; Kirschner, K. N.; Shields, G. C.; Hughes, J.; Lostritto, N.; Jacobson, H. I.; Andersen, T. T. Antiestrogenic and Anticancer Activities of Peptides Derived from the Active Site of Alpha-Fetoprotein. J. Pept. Sci. 2009, 15 (4), 319-325.
-
Salisburg, A. M.; Deline, A. L.; Lexa, K. W.; Shields, G. C.; Kirschner, K. N. Ramachandran-Type Plots for Glycosidic Linkages: Examples from Molecular Dynamic Simulations Using the Glycam06 Force Field. J. Comput. Chem. 2009, 30 (6), 910-921.
-
Shields, G. C. Computational Approaches for the Design of Peptides with Anti-Breast Cancer Properties. Future Med. Chem. 2009, 1 (1), 201-212.
-
Allodi, M. A.; Kirschner, K. N.; Shields, G. C. Thermodynamics of the Hydroxyl Radical Addition to Isoprene. J. Phys. Chem. A 2008, 112 (30), 7064-7071.
-
Dunn, M. E.; Shields, G. C.; Takahashi, K.; Skodje, R. T.; Vaida, V. Experimental and Theoretical Study of the OH Vibrational Spectra and Overtone Chemistry of Gas-Phase Vinylacetic Acid. J. Phys. Chem. A 2008, 112 (41), 10226-10235.
-
Hartt, G. M.; Shields, G. C.; Kirschner, K. N. Hydration of OCS with One to Four Water Molecules in Atmospheric and Laboratory Conditions. J. Phys. Chem. A 2008, 112 (19), 4490-4495.
-
Sherer, E. C.; Kirschner, K. N.; Pickard, F. C.; Rein, C.; Feldgus, S.; Shields, G. C. Efficient and Accurate Characterization of the Bergman Cyclization for Several Enediynes Including an Expanded Substructure of Esperamicin A(1). J. Phys. Chem. B 2008, 112 (51), 16917-16934.
-
Shields, G. C.; Kirschner, K. N. The Limitations of Certain Density Functionals in Modeling Neutral Water Clusters. Synth. React. Inorganic, Met. Nano-Metal Chem. 2008, 38 (1), 32-36.
-
Kirschner, K. N.; Hartt, G. M.; Evans, T. M.; Shields, G. C. In Search of CS2(H2O)n=1-4 Clusters. J. Chem. Phys. 2007, 126 (15).
-
Kirschner, K. N.; Lexa, K. W.; Salisburg, A. M.; Alser, K. A.; Joseph, L. C.; Andersen, T. T.; Bennett, J. A.; Jacobson, H. I.; Shields, G. C. Computational Design and Experimental Discovery of an Antiestrogenic Peptide Derived from Alpha-Fetoprotein. J. Am. Chem. Soc. 2007, 129 (19), 6263-6268.
-
Lexa, K. W.; Alser, K. A.; Salisburg, A. M.; Ellens, D. J.; Hernandez, L.; Bono, S. J.; Michael, H. C.; Derby, J. R.; Skiba, J. G.; Feldgus, S.; et al. The Search for Low Energy Conformational Families of Small Peptides: Searching for Active Conformations of Small Peptides in the Absence of a Known Receptor. Int. J. Quantum Chem. 2007, 107 (15), 3001-3012.
-
Allodi, M. A.; Dunn, M. E.; Livada, J.; Kirschner, K. N.; Shields, G. C. Do Hydroxyl Radical-Water Clusters, OH(H2O)n, n=1-5, Exist in the Atmosphere? J. Phys. Chem. A 2006, 110 (49), 13283-13289.
-
Alongi, K. S.; Dibble, T. S.; Shields, G. C.; Kirschner, K. N. Exploration of the Potential Energy Surfaces, Prediction of Atmospheric Concentrations, and Prediction of Vibrational Spectra for the HO2···(H2O)n (n = 1-2) Hydrogen Bonded Complexes. J. Phys. Chem. A 2006, 110 (10), 3686-3691.
-
Dunn, M. E.; Evans, T. M.; Kirschner, K. N.; Shields, G. C. Prediction of Accurate Anharmonic Experimental Vibrational Frequencies for Water Clusters, (H2O)n, n=2-5. J. Phys. Chem. A 2006, 110 (1), 303-309.
-
Pickard, F. C.; Griffith, D. R.; Ferrara, S. J.; Liptak, M. D.; Kirschner, K. N.; Shields, G. C. CCSD(T), W1, and Other Model Chemistry Predictions for Gas-Phase Deprotonation Reactions. Int. J. Quantum Chem. 2006, 106 (15, SI), 3122-3128.
-
Pickard, F. C.; Shepherd, R. L.; Gillis, A. E.; Dunn, M. E.; Feldgus, S.; Kirschner, K. N.; Shields, G. C.; Manoharan, M.; Alabugin, I. V. Ortho Effect in the Bergman Cyclization: Electronic and Steric Effects in Hydrogen Abstraction by 1-Substituted Naphthalene 5,8-Diradicals. J. Phys. Chem. A 2006, 110 (7), 2517-2526.
-
Day, M. B.; Kirschner, K. N.; Shields, G. C. Global Search for Minimum Energy (H2O)n Clusters, n=3-5. J. Phys. Chem. A 2005, 109 (30), 6773-6778.
-
Day, M. B.; Kirschner, K. N.; Shields, G. C. Pople-s Gaussian-3 Model Chemistry Applied to an Investigation of (H2O)8 Water Clusters. Int. J. Quantum Chem. 2005, 102 (5), 565-572.
-
Liptak, M. D.; Shields, G. C. Comparison of Density Functional Theory Predictions of Gas-Phase Deprotonation Data. Int. J. Quantum Chem. 2005, 105 (6), 580-587.
-
Pickard, F. C.; Dunn, M. E.; Shields, G. C. Comparison of Model Chemistry and Density Functional Theory Thermochemical Predictions with Experiment for Formation of Ionic Clusters of the Ammonium Cation Complexed with Water and Ammonia; Atmospheric Implications. J. Phys. Chem. A 2005, 109 (22), 4905-4910.
-
Pickard, F. C.; Pokon, E. K.; Liptak, M. D.; Shields, G. C. Comparison of CBS-QB3, CBS-APNO, G2, and G3 Thermochemical Predictions with Experiment for Formation of Ionic Clusters of Hydronium and Hydroxide Ions Complexed with Water. J. Chem. Phys. 2005, 122 (2).
-
Zhan, C. G.; Deng, S. X.; Skiba, J. G.; Hayes, B. A.; Tschampel, S. M.; Shields, G. C.; Landry, D. W. First-Principle Studies of Intermolecular and Intramolecular Catalysis of Protonated Cocaine. J. Comput. Chem. 2005, 26 (10), 980-986.
-
Dunn, M. E.; Pokon, E. K.; Shields, G. C. The Ability of the Gaussian-2, Gaussian-3, Complete Basis Set-QB3, and Complete Basis Set-APNO Model Chemistries to Model the Geometries of Small Water Clusters. Int. J. Quantum Chem. 2004, 100 (6), 1065-1070.
-
Dunn, M. E.; Pokon, E. K.; Shields, G. C. Thermodynamics of Forming Water Clusters at Various Temperatures and Pressures by Gaussian-2, Gaussian-3, Complete Basis Set-QB3, and Complete Basis Set-APNO Model Chemistries; Implications for Atmospheric Chemistry. J. Am. Chem. Soc. 2004, 126 (8), 2647-2653.
-
Palascak, M. W.; Shields, G. C. Accurate Experimental Values for the Free Energies of Hydration of H+, OH-, and H3O+. J. Phys. Chem. A 2004, 108 (16), 3692-3694.
-
Liptak, M. D.; Gross, K. C.; Seybold, P. G.; Feldgus, S.; Shields, G. C. Absolute pKa Determinations for Substituted Phenols. J. Am. Chem. Soc. 2002, 124 (22), 6421-6427.
-
Shields, G. C. The Benefits of Forming a Consortium for an NSF-MRI Proposal. CUR Q. 2002, December (December), 80-81.
-
Feldgus, S.; Shields, G. C. An ONIOM Study of the Bergman Reaction: A Computationally Efficient and Accurate Method for Modeling the Enediyne Anticancer Antibiotics. Chem. Phys. Lett. 2001, 347 (4-6), 505-511.
-
Liptak, M. D.; Shields, G. C. Accurate pKa Calculations for Carboxylic Acids Using Complete Basis Set and Gaussian-N Models Combined with CPCM Continuum Solvation Methods. J. Am. Chem. Soc. 2001, 123 (30), 7314-7319.
-
Liptak, M. D.; Shields, G. C. Experimentation with Different Thermodynamic Cycles Used for pKa Calculations on Carboxylic Acids Using Complete Basis Set and Gaussian-N Models Combined with CPCM Continuum Solvation Methods. Int. J. Quantum Chem. 2001, 85 (6, SI), 727-741.
-
Pokon, E. K.; Liptak, M. D.; Feldgus, S.; Shields, G. C. Comparison of CBS-QB3, CBS-APNO, and G3 Predictions of Gas Phase Deprotonation Data. J. Phys. Chem. A 2001, 105 (45), 10483-10487.
-
Sherer, E. C.; Bono, S. J.; Shields, G. C. Further Quantum Mechanical Evidence That Difluorotoluene Does Not Hydrogen Bond. J. Phys. Chem. B 2001, 105 (35), 8445-8451.
-
Toth, A. M.; Liptak, M. D.; Phillips, D. L.; Shields, G. C. Accurate Relative pKa Calculations for Carboxylic Acids Using Complete Basis Set and Gaussian-N Models Combined with Continuum Solvation Methods. J. Chem. Phys. 2001, 114 (10), 4595-4606.
-
Shields, G. C.; Laughton, C. A.; Orozco, M. Molecular Dynamics Simulation of a PNA·DNA·PNA Triple Helix in Aqueous Solution. J. Am. Chem. Soc. 1998, 120 (24), 5895-5904.
-
Sherer, E. C.; Yang, G.; Turner, G. M.; Shields, G. C.; Landry, D. W. Comparison of Experimental and Theoretical Structures of a Transition State Analogue Used for the Induction of Anti-Cocaine Catalytic Antibodies. J. Phys. Chem. A 1997, 101 (45), 8526-8529.
-
Shields, G. C.; Laughton, C. A.; Orozco, M. Molecular Dynamics Simulations of the d(T·A·T) Triple Helix. J. Am. Chem. Soc. 1997, 119 (32), 7463-7469.
-
Kirschner, K. N.; Sherer, E. C.; Shields, G. C. Use of the Supermolecule Approach to Model the Syn and Anti Conformations of Solvated Cyclic 3',5'-adenosine Monophosphate. J. Phys. Chem. 1996, 100 (8), 3293-3298.
-
Kirschner, K. N.; Shields, G. C. Quantum Mechanical Investigation of Cyclic 3',5'-adenosine Monophosphate, the Second Hormonal Messenger. J. Mol. Struct. THEOCHEM 1996, 362 (3), 297-304.
-
Sherer, E. C.; Turner, G. M.; Lively, T. N.; Landry, D. W.; Shields, G. C. A Semiempirical Transition State Structure for the First Step in the Alkaline Hydrolysis of Cocaine. Comparison between the Transition State Structure, the Phosphonate Monoester Transition State Analog, and a Newly Designed Thiophosphonate Transition Stat. J. Mol. Model. 1996, 2 (4), 62-69.
-
Brummel, H. A.; Shields, G. C. Semiempirical Study of the Bergman Reaction: Towards a Computationally Efficient and Accurate Method for Modeling Enediyne Anticancer Antibiotics. Int. J. Quantum Chem. 1995, No. 22, 51-59.
-
Sherer, E. C.; Turner, G. M.; Shields, G. C. Investigation of the Potential Energy Surface for the First Step in the Alkaline Hydrolysis of Methyl Acetate. Int. J. Quantum Chem. 1995, No. 22, 83-93.
-
Turner, G. M.; Sherer, E. C.; Shields, G. C. A Computationally Efficient Procedure for Modeling the First Step in the Alkaline Hydrolysis of Esters. Int. J. Quantum Chem. 1995, No. 22, 103-112.
-
Kash, M. M.; Shields, G. C. Using the Franck-Hertz Experiment to Illustrate Quantization: Energy States of the Neon Atom by Electron Impact. J. Chem. Educ. 1994, 71 (6), 466-468.
-
Kirschner, K. N.; Shields, G. C. Quantum-Mechanical Investigation of Large Water Clusters. Int. J. Quantum Chem. 1994, No. 28, 349-360.
-
Lively, T. N.; Jurema, M. W.; Shields, G. C. Hydrogen Bonding of Nucleotide Base Pairs: Application of the PM3 Method. Int. J. Quantum Chem. 1994, No. 21, 95-107.
-
Shields, G. C. The Physical Chemistry Sequence at Liberal Arts Colleges: The Lake Forest College Approach. J. Chem. Educ. 1994, 71 (11), 951-953.
-
Jurema, M. W.; Kirschner, K. N.; Shields, G. C. Modeling of Magic Water Clusters (H2O)20 and (H2O)21H+ with the PM3 Quantum-Mechanical Method. J. Comput. Chem. 1993, 14 (11), 1326-1332.
-
Jurema, M. W.; Shields, G. C. Ability of the PM3 Quantum-Mechanical Method to Model Intermolecular Hydrogen Bonding between Neutral Molecules. J. Comput. Chem. 1993, 14 (1), 89-104.
-
Shields, G. C.; Kash, M. M. Experiment in Quantization: Atomic Line Spectra. J. Chem. Educ. 1992, 69 (4), 329-331.
-
Kalkanis, G. H.; Shields, G. C. AM1 and PM3 Calculations of the Potential Energy Surfaces for Hydroxymethyl Radical Reactions with Nitric Oxide and Nitrogen Dioxide. J. Phys. Chem. 1991, 95 (13), 5085-5089.
-
Schultz, S. C.; Shields, G. C.; Steitz, T. A. Crystal Structure of a CAP-DNA Complex: The DNA Is Bent by 90 Degrees. Science 1991, 253 (5023), 1001-1007.
-
Schultz, S. C.; Shields, G. C.; Steitz, T. A. Crystallization of Escherichia coli Catabolite Gene Activator Protein with Its DNA Binding Site: The Use of Modular DNA. J. Mol. Biol. 1990, 213 (1), 159-166.
-
Nelson, P. R.; Fung, C.; SedgwickK, J. B.; Shields, G. C.; Abbey, L. E.; Moran, T. F. Doubly Charged Ion Mass Spectra of Alkyl-Substituted Furans and Pyrroles. Org. Mass Spectrom. 1987, 22 (7), 389-399.
-
SedgwickK, J. B.; Paulson, B. P.; Shields, G. C.; Moran, T. F. Competition between Single and Double Electron Transfer in Collisions of Doubly Charged Molecular Pyrrole Ions with Neutral Pyrrole Molecules. Int. J. Mass Spectrom. Ion Process. 1987, 79 (1), 127-140.
-
Shields, G. C.; Steiner, P. A.; Nelson, P. R.; Trauner, M. C.; Moran, T. F. Charge Transfer Reactions of Organic Ions Containing Oxygen: Correlation between Reaction Energetics and Cross Sections. Org. Mass Spectrom. 1987, 22 (2), 64-69.
-
Steitz, T. A.; Beese, L.; Engelman, B.; Freemont, P.; Friedman, J.; Sanderson, M.; Schultz, S.; Shields, G.; Warwicker, J. Structural Studies of Three DNA Binding Proteins: Catabolite Gene Activator Protein, Resolvase, and the Klenow Fragment of DNA Polymerase I. In DNA-Ligand Interactions; Springer US: Boston, MA, 1987; pp 185-189.
-
Appling, J. R.; Shields, G. C.; Moran, T. F. Evidence for Long-Lived Excited States of [CnH2]2+ Carbodications. Org. Mass Spectrom. 1986, 21 (2), 69-75.
-
Burdick, G. W.; Shields, G. C.; Moran, T. F. Polarizabilites of Organic Ions. Org. Mass Spectrom. 1986, 21 (7), 449-450.
-
Shields, G. C.; Moran, T. F. Doubly-Charged Ethane Ions: Solution to the Dilemma of Stability Predicted by Theory and Instability Observed in Experiment. Org. Mass Spectrom. 1986, 21 (8), 479-483.
-
Shields, G. C.; Moran, T. F. Doubly-Charged Gas Phase Cations. Theor. Chim. Acta 1986, 69 (2), 147-159.
-
Shields, G. C.; Wennberg, L.; Wilcox, J. B.; Moran, T. F. Sensitivity of Charge Transfer Reactions to Hydrocarbon Ion Structures. Org. Mass Spectrom. 1986, 21 (3), 137-149.
-
Burdick, G. W.; Shields, G. C.; Appling, J. R.; Moran, T. F. Structures, Energetics and Fragmentation Pathways of CnH22+ Carbodications. Int. J. Mass Spectrom. Ion Process. 1985, 64 (3), 315-333.
-
Shields, G. C.; Moran, T. F. Molecular Charge-Transfer Cross Sections and Their Correlation with Reactant Ion Structures. J. Phys. Chem. 1985, 89 (19), 4027-4031.
-
Shields, G. C.; Moran, T. F. Double Electron Transfer Reactions of CO22+ Ions. Chem. Phys. Lett. 1983, 101 (3), 287-290.
-
Shields, G. C.; Moran, T. F. Single- and Double-Electron Transfer Reactions of Ground and Metastable State Ar2+ Ions. J. Phys. B - At. Mol. Opt. Phys. 1983, 16 (19), 3591-3601.