Fungi focus of student’s simulation
Just like bacteria, fungi can develop resistance to drugs used to treat infections, a “growing threat” across the country, according to the Centers for Disease Control and Prevention. With only three classes of anti-fungal medications, the need to find new ways to combat fungal infections is increasingly important.
One Furman University student is contributing to the understanding of drug-resistant fungi. Mason Dudley ’23 is working to create a computer model of a protein that causes drug resistance in the fungus Candida glabrata. He’s working with chemistry professor George Shields, chemistry assistant professor Meghan Breen and Karl Kirschner, who is on faculty at Bonn Rhein-Sieg University of Applied Sciences in Germany.
We asked Dudley about his research. His answers are below.
What’s your research about?
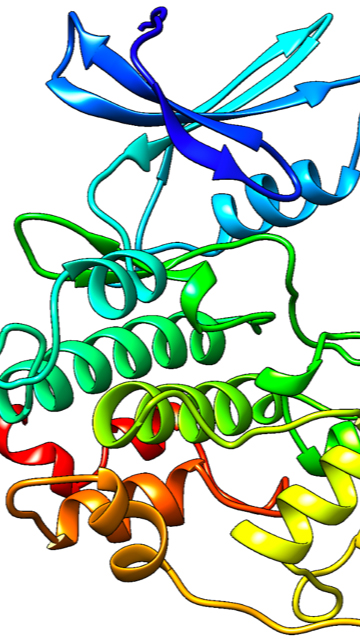
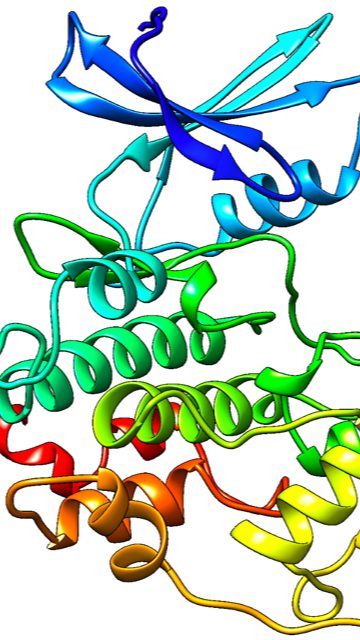
We are working creating a model of a protein, Pho85, that is responsible for drug resistance in C. glabrata, a common fungal infection, and we are also working on simulating the activation of it as well. A cyclin is a protein that has roles in cell division, in this case when our protein Pho85 is paired with a cyclin it has additional rolls in the cell such as the phosphorous pathway, which is why we are interested in this protein.
My research is focused on computationally investigating a cyclin dependent kinase (CDK) that, when phosphorylated, is responsible for activation of PDR1, a transcription factor that causes azole drug resistance in C. glabrata. So far, I modeled the protein as well as its major cyclin, Pho80, and have begun to run molecular dynamics to understand how this protein acts within the body.
Why does this research interest you?
This research is interesting as azoles are becoming a less effective treatment for fungal infections, especially with C. glabrata, leading to a possible rise in super fungi.
Where has your research taken you so far?
We have generated a visual that shows the flexibility of the cdk itself, as well shows the motion of the ATP binding in Pho85. I am currently working on using a mix of quantum mechanics and molecular mechanics to simulate the hydrolysis of ATP within this protein, where a single phosphate will transfer to the protein simulating the theoretical mechanism of activation leading to activation of PDR1. I have a video that shows the movement of protein throughout the molecular dynamics simulations if you want me to send it to you.
Has anything surprised you?
Not much has surprised me other than how fast and how much data I can generate. One 100 nano-second simulation of this protein generates a 100-gigabyte trajectory file that helps us to visualize the movement. So far, I have generated over 10 terabytes of data for this protein, having made 100 replicas, and have plans to generate a total of over 50TB of data when I begin to run molecular dynamics on its cyclin Pho80.
What does being able to do research mean to you?
Research to me means being to learn many different skills and work with a diverse group of people.